Statistical and computational methods
1. Single cell genomics
Single-cell genomic technologies such as single-cell RNA-seq and single-cell ATAC-seq provide unprecedented power for examining the functional genomic landscape of a heterogeneous cell population. We develop statistical and computational methods and tools for designing single-cell genomic experiments and analyzing single-cell genomic data. Examples of our tools include TSCAN, SCATE, BIRD, SCRAT, Lamian and TreeCorTreat.
2. High-throughput regulome and epigenome profiling and analysis
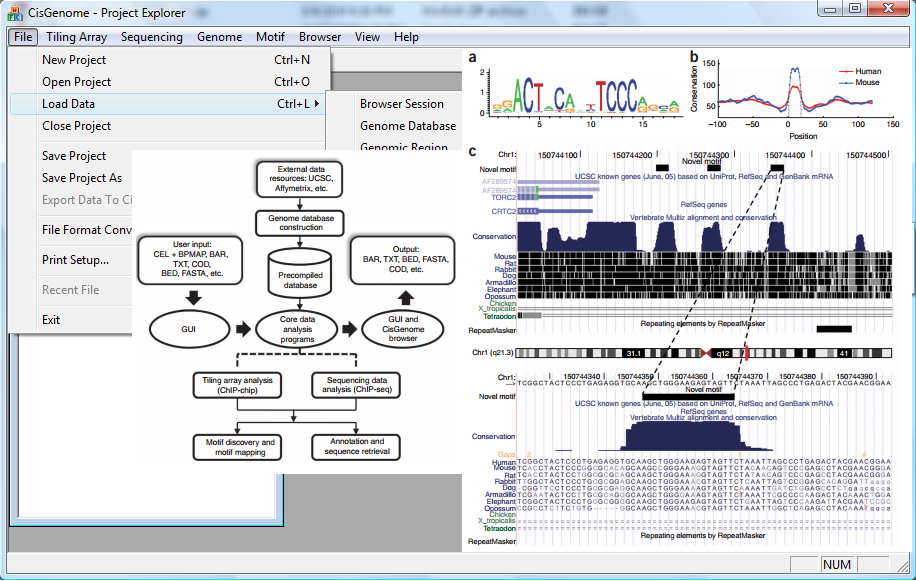
Regulome and epigenome provide key information to understand gene regulation. We develop analytical and software tools for analyzing regulome and epigenome data generated by high-throughput technologies such as ChIP-seq, DNase-seq, ATAC-seq, etc. Examples of our tools include CisGenome, dPCA, TileMap, TileProbe, and JAMIE. We have also developed a database, hmChIP, to help scientists to explore publicly available ChIP-seq and ChIP-chip data.
3. High-throughput transcriptome analysis and integration
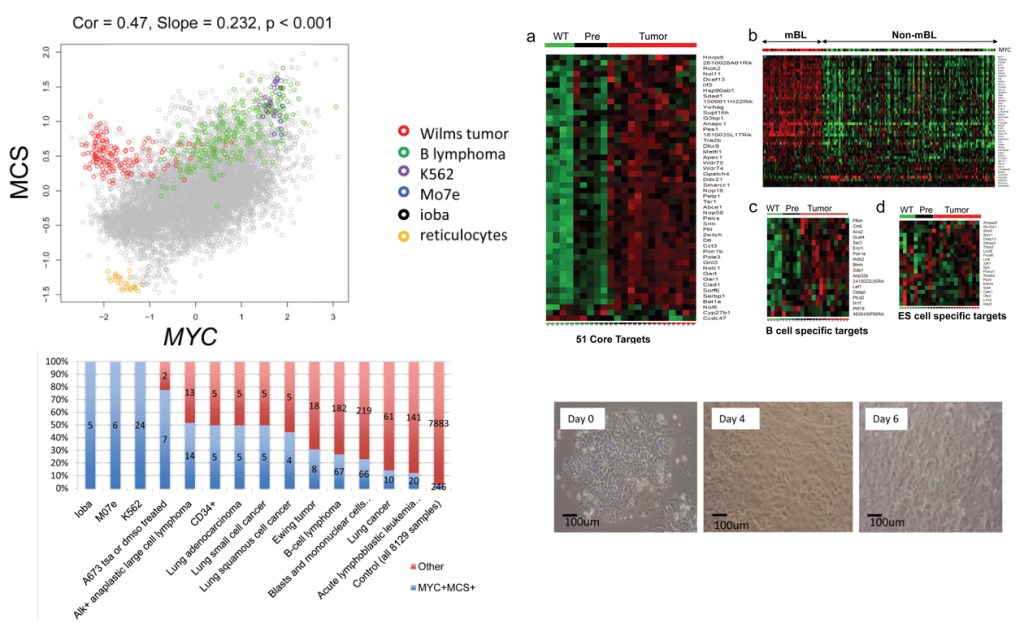
We develop methods for analyzing large scale gene expression data. One example is the correlation motif approach, CorMotif, for integrative analysis of multiple gene expression experiments. Another example is Gene Set Context Analysis (GSCA), a method to help researchers systematically identify cell types, conditions and diseases associated with user-specified gene set activity patterns.
4. Sequence motif discovery and analysis
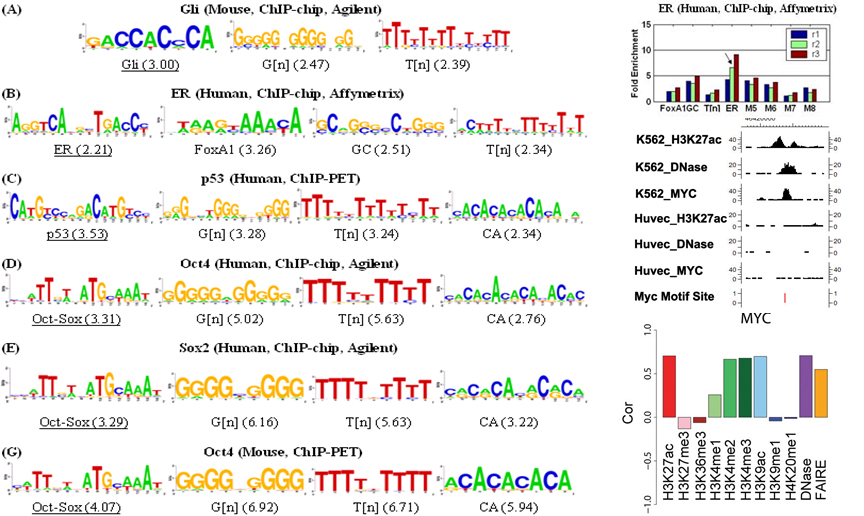
5. Scalable data integration
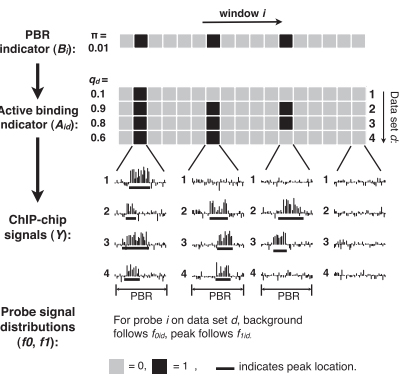
Integrative ‘omics analysis can lead to new discoveries. Data integration and data mining are non-trivial. Common issues include high dimensionality, heterogeneity, complex correlation structure, exponential computation complexity, etc. We develop methods and tools for data integration that tackle these challenges. Examples include BIRD, a big data regression method for predicting genome-wide regulatory element activities using gene expression, iASeq for integrative analysis of allele-specificity, JAMIE for joint analysis of multiple ChIP-chip datasets, CorMotif for joint analysis of multiple gene expression datasets, ChIP-PED for joint analysis of ChIP and public gene expression data.
Applications to biology, medicine and public health
6. Decoding gene regulation in stem cells, development and diseases
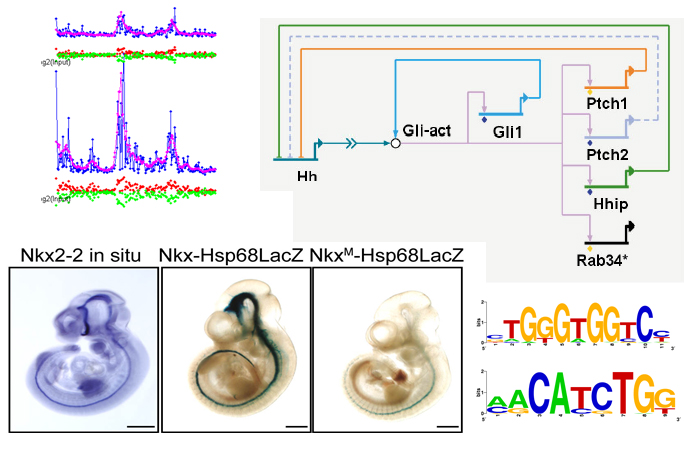
Gene activities are tightly controlled both temporally and spatially. We are interested in decoding gene regulatory programs in development, stem cells and diseases. We have contributed to understanding gene regulation in a variety of systems. Examples include (1) human and mouse embryonic stem cells [1,2], (2) the sonic hedgehog signaling pathway in embryonic development [3,4,5], (3) B cell lymphoma [1], leukemia [6], and various other cancers [7].
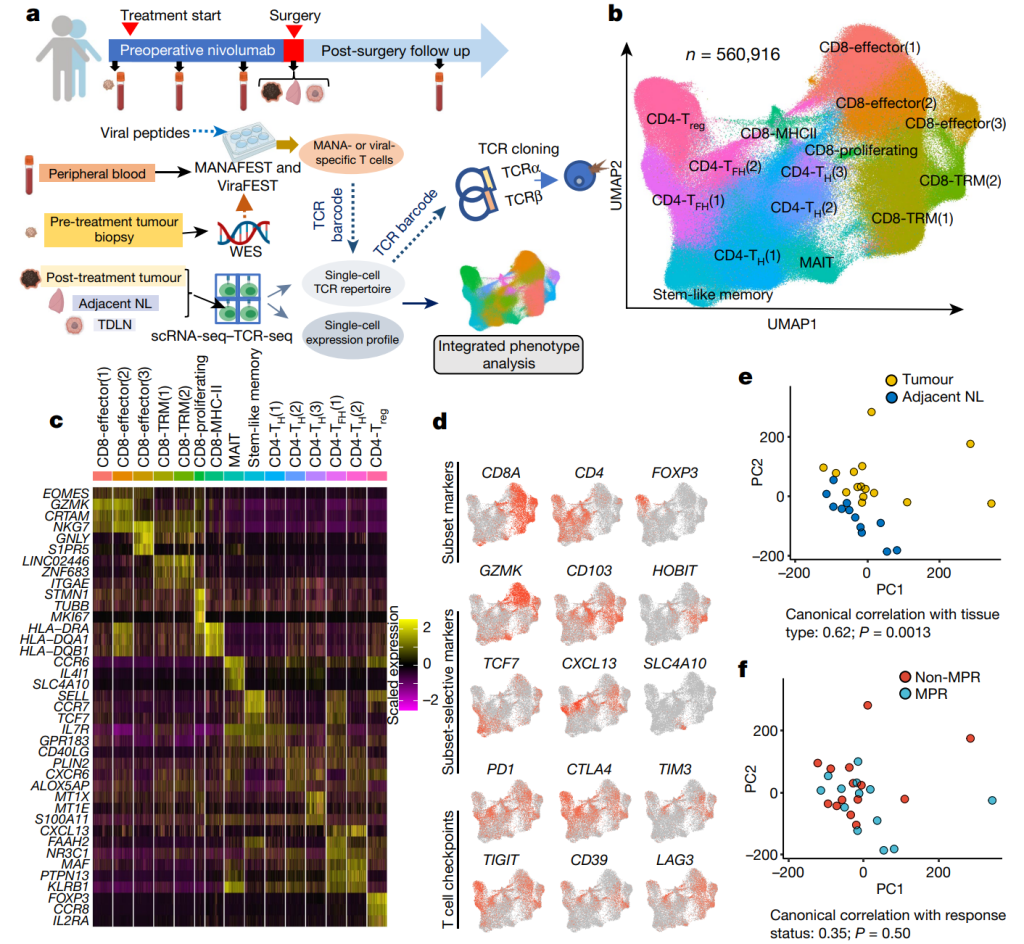
7. Immunology in cancer and infectious disease
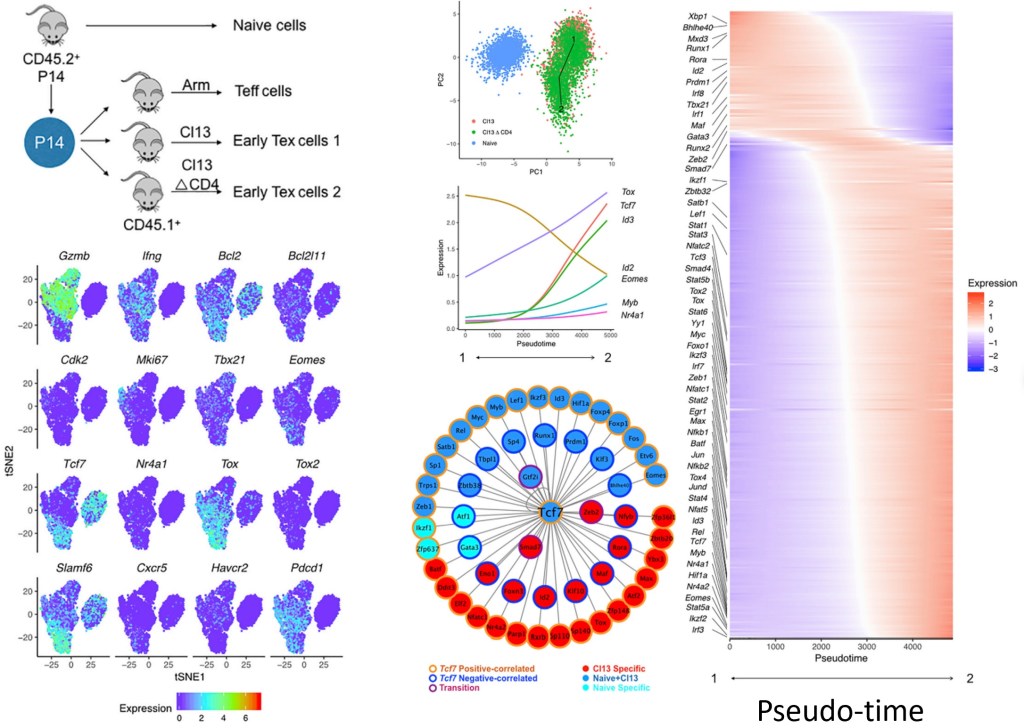
Understanding the immune system is crucial to understand how our bodies respond to viral infection, tumor antigens, immunotherapy, and vaccines, etc. We develop methods and tools that analyze single-cell genomic and immune profiling data and we use these tools to study how our immune system works in cancer and infectious diseases. [1,2,3]